Performing a permutation test
Overview
Teaching: 10 min
Exercises: 20 minQuestions
How can I evaluate the statistical significance of genome scan results?
Objectives
Run a permutation test to establish LOD score thresholds.
To establish the statistical significance of the results of a genome scan, a permutation test can identify the maximum LOD score that can occur by random chance. A permutation tests shuffles genotypes and phenotypes, essentially breaking the relationship between the two. The genome-wide maximum LOD score is then calculated on the permuted data, and this score used as a threshold of statistical significance. A genome-wide maximum LOD on shuffled, or permuted, data serves as the threshold because it represents the highest LOD score generated by random chance.
The scan1perm() function takes the same arguments as scan1(), plus additional arguments to control the permutations:
n_permis the number of permutation replicates.perm_Xspcontrols whether to perform autosome/X chromosome specific permutations (withperm_Xsp=TRUE) or not (the default is FALSE).perm_stratais a vector that defines the strata for a stratified permutation test.chr_lengthsis a vector of chromosome lengths, used in the case thatperm_Xsp=TRUE.
As with scan1(), you may provide a kinship matrix (or vector of kinship matrices, for the “leave one chromosome out” (loco) approach), in order to fit linear mixed models. If kinship is unspecified, the function performs ordinary Haley-Knott regression.
To perform a permutation test with the iron data, we run scan1perm(), provide it with the genotype probabilities, the phenotype data, X covariates and number of permutations. For expediency, we’ll use only 10 permutations, although 1000 is recommended.
operm <- scan1perm(genoprobs = pr, pheno = iron$pheno, Xcovar = Xcovar, n_perm = 1000) # replace 1000 with 10 for expediency
Note the need to specify special covariates for the X chromosome (via Xcovar), to be included under the null hypothesis of no QTL. And note that when these are provided, the default is to perform a stratified permutation test, using strata defined by the rows in
Xcovar. In general, when the X chromosome is considered, one will wish to stratify at least by sex.
Also note that, as with scan1(), you can speed up the calculations on a multi-core machine by specifying the argument cores. With cores=0, the number of available cores will be detected via parallel::detectCores(). Otherwise, specify the number of cores as a positive integer. For large datasets, be mindful of the amount of memory that will be needed; you may need to use fewer than the maximum number of cores, to avoid going beyond the available memory.
operm <- scan1perm(pr, iron$pheno, Xcovar=Xcovar, n_perm=1000, cores=0)
operm now contains the maximum LOD score for each permutation for the liver and spleen phenotypes. There should be 1000 values for each phenotypes. We can view the liver permutation LOD scores by making a histogram.
hist(operm[,'liver'], breaks = 50, xlab = "LOD", main = "LOD scores for liver scan with threshold in red")
abline(v = summary(operm)[,'liver'], col = 'red', lwd = 2)
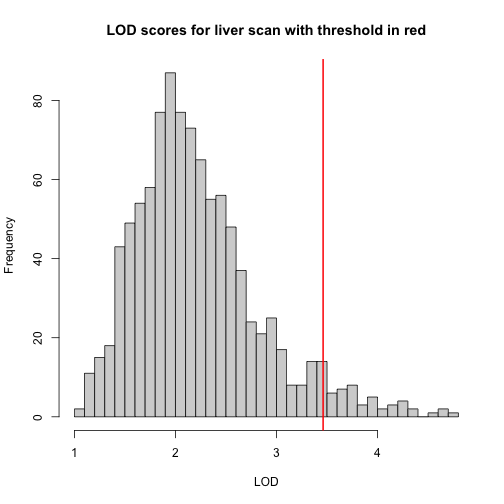
In the histogram above, you can see that most of the maximum LOD scores fall between 1 and 3. This means that we expect LOD scores less than 3 to occur by chance fairly often. The red line indicates the alpha = 0.05 threshold, which means that, under permutation, we only see LOD values as high or higher, 5% of the time. This is one way of estimating a significance threshold for QTL plots.
To get estimated significance thresholds, use the function summary().
summary(operm)
LOD thresholds (1000 permutations)
liver spleen
0.05 3.34 3.79
The default is to return the 5% significance thresholds. Thresholds for other (or for multiple) significance levels can be obtained via the alpha argument.
summary(operm, alpha=c(0.2, 0.05))
LOD thresholds (1000 permutations)
liver spleen
0.2 2.65 2.73
0.05 3.34 3.79
To obtain autosome/X chromosome-specific significance thresholds, specify perm_Xsp=TRUE. In this case, you need to provide chromosome lengths, which may be obtained with the function chr_lengths().
operm2 <- scan1perm(pr, iron$pheno, Xcovar=Xcovar, n_perm=1000,
perm_Xsp=TRUE, chr_lengths=chr_lengths(map))
Separate permutations are performed for the autosomes and X chromosome, and considerably more permutation replicates are needed for the X chromosome. The computations take about twice as much time. See Broman et al. (2006) Genetics 174:2151-2158.
The significance thresholds are again derived via summary():
summary(operm2, alpha=c(0.2, 0.05))
Autosome LOD thresholds (1000 permutations)
liver spleen
0.2 2.64 2.62
0.05 3.33 3.32
X chromosome LOD thresholds (28243 permutations)
liver spleen
0.2 3.13 4.01
0.05 3.74 5.03
As with scan1, we can use scan1perm with binary traits, using the argument model="binary". Again, this can’t be used with a kinship matrix, but all of the other arguments can be applied.
operm_bin <- scan1perm(pr, bin_pheno, Xcovar=Xcovar, n_perm=1000,
perm_Xsp=TRUE, chr_lengths=chr_lengths(map),
model="binary")
Here are the estimated 5% and 20% significance thresholds.
summary(operm_bin, alpha=c(0.2, 0.05))
Autosome LOD thresholds (1000 permutations)
liver spleen
0.2 2.68 2.61
0.05 3.45 3.33
X chromosome LOD thresholds (28243 permutations)
liver spleen
0.2 3.09 3.12
0.05 3.80 3.81
The code below shuffles the phenotypes so that they no longer match up with the genotypes. The purpose of this is to find out how high the LOD score can be due to random chance alone.
shuffled_order <- sample(rownames(iron$pheno))
pheno_permuted <- iron$pheno
rownames(pheno_permuted) <- shuffled_order
xcovar_permuted <- Xcovar
rownames(xcovar_permuted) <- shuffled_order
out_permuted <- scan1(genoprobs = pr, pheno = pheno_permuted, Xcovar = xcovar_permuted)
plot(out_permuted, map)
head(shuffled_order)
Challenge 1
Run the preceding code to shuffle the phenotype data and plot a genome scan with this shuffled (permuted) data.
What is the maximum LOD score in the scan from this permuted data?
How does it compare to the maximum LOD scores obtained from the earlier scan?
How does it compare to the 5% and 20% LOD thresholds obtained earlier?
Paste the maximum LOD score in the scan from your permuted data into the etherpad.Solution to Challenge 1
Challenge 2
1) Find the 1% and 10% significance thresholds for the first set of permutations contained in the object
operm.
2) What do the 1% and 10% significance thresholds say about LOD scores?Solution to Challenge 2
1)
summary(operm, alpha=c(0.01, 0.10))
2) These LOD thresholds indicate maximum LOD scores that can be obtained by random chance at the 1% and 10% significance levels. We expect to see LOD values this high or higher 1% and 10% of the time respectively.
Key Points
A permutation test establishes the statistical significance of a genome scan.